guest post by Tom Leinster
Even if there weren’t a global biodiversity crisis, we’d want to know how to put a number on biodiversity. As Lord Kelvin famously put it:
When you can measure what you are speaking about, and express it in numbers, you know something about it; but when you cannot measure it, your knowledge is of a meagre and unsatisfactory kind: it may be the beginning of knowledge, but you have scarcely, in your thoughts, advanced to the stage of science.
In this post, I’ll talk about what happens when you take a mass of biological data and try to turn it into a single number, intended to measure biodiversity.
There have been more than 50 years of debate about how to measure diversity. While the idea of putting a number on biological diversity goes back to the 1940s at least, the debate really seems to have got going in the wake of pioneering work by the great ecologist Robert Whittaker in the 1960s.
There followed several decades in which progress was made… but there was a lot of talking at cross-purposes. In fact, there was so much confusion that some people gave up on the diversity concept altogether. The mood is summed up by the title of an excellent and much-cited paper of Stuart Hurlbert:
• S. H. Hurlbert, The nonconcept of species diversity: A critique and alternative parameters. Ecology 52:577–586, 1971.
So why all the confusion?
One reason is that the word “diversity” is used by different people in many different ways. We all know that diversity is important: so if you found a quantity that seemed to measure biological variation in a sensible way, you might be tempted to call it “diversity” and publish a paper promoting your quantity over all other quantities that have ever been given that name. There are literally dozens of measures of diversity in the literature. Here are two simple ones:
- Species richness is simply the number of species in the community concerned.
-
The Shannon entropy is
, where our community consists of
species in proportions
.
Which quantity should we call “diversity”? Do all these quantities really measure the same kind of thing? If community A has greater than species richness than community B, but lower Shannon entropy, what does it mean?
Another cause for confusion is a blurring between the questions
Which quantities deserve to be called diversity?
and
Which quantities are we capable of measuring experimentally?
For example, we might all agree that species richness is an important quantity, but that doesn’t mean that species richness is easy to measure in practice. (In fact, it’s not, more on which below.) My own view is that the two questions should be kept separate:
The statistical problem of designing appropriate estimators becomes relevant only after the measure to be estimated is accepted to be meaningful.
(Hans-Rolf Gregorius, Elizabeth M. Gillet, Generalized Simpson-diversity, Ecological Modelling 211:90–96, 2008.)
The problems involved in quantifying diversity are of three types: practical, statistical and conceptual. I’ll say a little about the first two, and rather more about the third.
Practical Suppose that you’re doing a survey of the vertebrates in a forest. Perhaps one important species is brightly coloured and noisy, while another is silent, shy, and well-camouflaged. How do you prevent the first from being recorded disproportionately?
Or suppose that you’re carrying out a survey, with multiple people doing the fieldwork. Different people have a tendency to spot different things: for example, one person might be short-sighted and another long-sighted. How do you ensure that this doesn’t affect your results?
Statistical Imagine that you want to know how many distinct species of insect live in a particular area — the “species richness”, in the terminology introduced above. You go out collecting, and you come back with 100 specimens representing 10 species.
But your survey might have missed some species altogether, so you go out and get a bigger sample. This time, you get 200 specimens representing 15 species. Does this help you discover how many species there really are?
Logically, not at all. The only certainty is that there are at least 15 species. Maybe there are thousands of species, but almost all of them are extremely rare. Or maybe there are really only 15. Unless you collect all the insects, you’ll never know for sure exactly how many species there are.
However, it may be that you can make reasonable assumptions about the frequency distribution of the species. People sometimes do exactly this, to try to overcome the difficulty of estimating species richness.
Conceptual This is what I really want to talk about.
I mentioned earlier that different people mean different things by “diversity”. Here’s an example.
Consider two bird communities. The first looks like this:

It contains four species, one of which is responsible for most of the population, and three of which are quite rare. The second looks like this:

It has only three species, but they’re evenly balanced.
Which community is the more diverse? It’s a matter of opinion. Mostly in the press, and in many scholarly articles too, “biodiversity” is used as a synonym for “species richness”. On this count, the first community is more diverse. But if you’re more concerned with the healthy functioning of the whole community, the presence of rare species might not be particularly important: it’s balance that matters, and the second community has more of that.
Different people using the word “diversity” attach different amounts of significance to rare species. There’s a spectrum of points of view, ranging from those who give rare species the same weight as common ones (as in the definition of species richness) to those who are only interested in the most common species of all. Every point on this spectrum of viewpoints is reasonable. None should have a monopoly on the word “diversity”.
At least, that’s what Christina Cobbold and I argue in our new paper:
• Tom Leinster, Christina A. Cobbold, Measuring diversity: the importance of species similarity, Ecology, in press (doi:10.1890/10-2402.1).
But that’s not actually our main point. As the title suggests, the real purpose of our paper is to show how to measure diversity in a way that reflects the varying differences between species. I’ll explain.
Most of the existing approaches to measuring biodiversity go like this.
We have a “community” of organisms — the fish in a lake, the fungi in a forest, or the bacteria on your skin. This community is divided into groups, conventionally called species, though they needn’t be species in the ordinary sense.
We assume that we know the relative abundances, or relative frequencies, of the species. Write them as . Thus,
is the proportion of the total population that belongs to the
th species, where “proportion” is measured in any way you think sensible (number of individuals, total mass, etc).
We only care about relative abundances here, not absolute abundances: so . If half of a forest is destroyed, it might be a catastrophe, but on the (unrealistic) assumption that all the flora and fauna in the forest were distributed homogeneously, it won’t actually change the biodiversity. (That’s not a statement about what’s important in life; it’s only a statement about the usage of a word.)
This model is common but crude. It can’t detect the difference between a community of six dramatically different species and a community consisting of six species of barnacle.
So, Christina and I use a refined model, as follows. We assume that we also have a measure of the similarity between each pair of species. This is a real number between 0 and 1, with 0 indicating that the species are as dissimilar as could be, and 1 indicating that they’re identical. Writing the similarity between the th and
th species as
, this gives an
matrix
. Our only assumption on
is that its diagonal entries are all 1: every species is identical to itself.
There are many ways of measuring inter-species similarity. Probably the most familiar approach is genetic, as in “you share 98% of your DNA with a chimpanzee”. But there are many other possibilities: functional, phylogenetic, morphological, taxonomic, …. Diversity is a measure of the variety of life; having to choose a measure of similarity forces you to get clear exactly what you mean by “variety”.
Christina and I are by no means the first people to incorporate species similarity into the model of an ecological community. The main new thing in our paper is this measure of the community’s diversity:
What does this mean?
-
is what we call the diversity of order
of the community. Here
is a parameter between
and
, which you get to choose. Different values of
represent different points on the spectrum of viewpoints described above. Small values of
give high importance to rare species; large values of
give high importance to common species.
-
is shorthand for the relative abundances
, and
is the matrix of similarities.
-
means
.
The expression doesn’t make sense if or
, but can be made sense of by taking limits. For
, this gives
If you want to know the value at , or any of the other mathematical details, you can read this post at the n-Category Café, or of course our paper. In both places, you’ll also find an explanation of what motivates this formula. What’s more, you’ll see that many existing measures of diversity are special cases of ours, obtained by taking particular values for
and/or
.
But I won’t talk about any of that here. Instead, I’ll tell you how taking species similarity into account can radically alter the assessment of diversity.
I’ll do this using an example: butterflies of subfamily Charaxinae at a site in an Ecuadorian rainforest. The data is from here:
• P. J. DeVries, D. Murray, R. Lande, Species diversity in vertical, horizontal and temporal dimensions of a fruit-feeding butterfly community in an Ecuadorian rainforest. Biological Journal of the Linnean Society 62:343–364, 1997.
They measured the butterfly abundances in both the canopy (top level) and understorey (lower level) at this site, with the following results:
| Species | Canopy | Understorey |
| Prepona laertes | 15 | 0 |
| Archaeoprepona demophon | 14 | 37 |
| Zaretis itys | 25 | 11 |
| Memphis arachne | 89 | 23 |
| Memphis offa | 21 | 3 |
| Memphis xenocles | 32 | 8 |
Which is more diverse: canopy or understorey?
We’ve already seen that the answer is going to depend on what exactly we mean by “diverse”.
First let’s answer the question under the (crude!) assumption that different species have nothing whatsoever in common. This means taking our similarity matrix to be the identity matrix: if
then
(totally dissimilar), and if
then
(totally identical).
Now, remember that there’s a spectrum of viewpoints on how much importance to give to rare species when measuring diversity. Rather than choosing a particular viewpoint, we’ll calculate the diversity from all viewpoints, and display it on a graph. In other words, we’ll draw the graph of (the diversity of order
) against
(the viewpoint). Here’s what we get:

(the horizontal axis should be labelled with a .)
Conclusion: from all viewpoints, the butterfly population in the canopy is at least as diverse as that in the understorey.
Now let’s do it again, but this time taking account of the varying similarities between species of butterflies. We don’t have much to go on: how do we know whether Prepona laertes is very similar to, or very different from, Archaeoprepona demophon? With only the data above, we don’t. So what can we do?
All we have to go on is the taxonomy. Remember your high school biology: for the butterfly Prepona laertes, the genus is Prepona and the species is laertes. We’d expect species in the same genus to have more in common than species in different genera. So let’s define the similarity between two species as follows:
- the similarity is 1 if the species are the same
- the similarity is 0.5 if the species are different but in the same genus
- the similarity is 0 if they are not even in the same genus.
This is still crude, but in the absence of further information, it’s about the best we can do. And it’s better than the first approach, where we ignored the taxonomy entirely. Throwing away biologically relevant information is unlikely to lead to a better assessment of diversity.
Using this taxonomic matrix , and the same abundances, the diversity graphs become:

This is more interesting! For , the understorey looks more diverse than the canopy — the opposite conclusion to our first approach.
It’s not hard to see why. Look again at the table of abundances, but paying attention to the genera of the butterflies. In the canopy, nearly three-quarters of the butterflies are of genus Memphis. So when we take into account the fact that species in the same genus tend to be somewhat similar, the canopy looks much less diverse than it did before. In the understorey, however, the species are spread more evenly between genera, so taking similarity into account leaves its diversity relatively unchanged.
Taking account of species similarity opens up a world of uncertainty. How should we measure similarity? There are as many possibilities as there are quantifiable characteristics of living organisms. It’s much more reassuring to stay in the black-and-white world where distinct species are always assigned a similarity of 0, no matter how similar they might actually be. (This is, effectively, what most existing measures do.) But that’s just hiding from reality.
Maybe you disagree! If so, try the the Discussion section of our paper, where we lay out our arguments in more detail. Or let me know by leaving a comment.

Interesting. Do you have a sense of how to measure diversities of ecosystems that take into account interactions between species? It is not unusual to read that this ecosystem is “simple” and this one is “complex.” Is there a way to quantify this?
For some reason I have a feeling that simplicial sets should come in.
Interactions are the difference (or part of the difference) between diversity and complexity. That’s one of the problems for complexity-stability research in ecology. Theoretical work tends to look at complexity, but empirical work focuses on diversity because it’s much more tractable.
Now I’m off to look up “simplicial set”.
Hello again, Jane!
I would love to learn more about ‘complexity’ in ecology. Since I come from a theoretical physics background, I like interactions: I like stuff to happen. Are there any good recent review articles? I just know some old classic arguments about the stability of food webs—my current (lack of) understanding is documented here:
• Systems ecology, Azimuth Library.
Nuances aside, you can roughly think of a simplicial set as a bunch of dots, a bunch of oriented edges, a bunch of triangles, a bunch of tetrahedra, and so on:
(There should be arrows on these edges.) If all the triangles and higher-dimensional simplexes are ‘degenerate’ (that is squashed down so you can’t see them), a simplicial set is just a directed graph.
One way to build a more interesting simplicial set is to take a bunch of species (say), use those as the vertices, and throw in a simplex with vertices (a,b,c,d,…) whenever all the species corresponding to those vertices are related (or interact) in some specified way. This only works if whenever a collection of species counts as related, all its sub-collections also count as related. Simplicial sets built this way show up in a lot of applications.
(I beg experts not to jump in right now pointing out the distinction between simplicial sets, simplical complexes, and symmetric sets. These details matter, but only after one thinks the subject is interesting!)
I’m also much more interested in ecological dynamics than static diversity. The place to start is with simple Lotka-Volterra models. They’re still used in some research and any population ecology or theoretical ecology book will have a lot to say about them.
The n-species Yodzis-Innes model incorporates more realistic assumptions about energetic constraints on parameters and how predator feeding rates change with prey abundance. See:
• Richard J. William, Homage to Yodzis and Innes 1992: scaling up feeding-based population dynamics to complex ecological networks.
Actually, much of the material at foodwebs.org is worth reading — the PeaceLab folks do a lot of very good work on food webs.
For a good overview of diversity, complexity and stability, see:
• James Justus, Complexity, diversity and stability.
Yes, Justus is in a philosophy department, but most of that chapter is just a good explanation of key ecological concepts — I’d happily assign it to grad students.
Jane wrote:
There’s obviously much more richness to be found in a dynamic situation than a merely static one. Nevertheless, the relatively simple problem of quantifying the diversity of a single static community shouldn’t be underestimated: it’s been tripping people up for decades.
Justus, in the book chapter you link to, cites a 2005 paper of Ricotta called “Through the jungle of biological diversity”. The thrust of that paper, as I understood it, is that there’s a bewildering mess of diversity measures and it’s near-impossible to hack your way through them. This seems to have been a common view at the time, and maybe in some circles it still is.
But since 2006 (possibly before Justus wrote that chapter), things have become much clearer. The turning point was a paper of Lou Jost arguing that all diversity measures should be effective numbers, meaning that the diversity value assigned to a community of species in equal proportions is
species in equal proportions is  . There are several excellent reasons for this; see the paper for some of them.
. There are several excellent reasons for this; see the paper for some of them.
As Jost showed, you can transform any diversity index into an effective number (its “numbers equivalent”) by a few simple steps of algebra. Let’s do this for two of the measures mentioned in Justus’s chapter. First, the Gini-Simpson index
gets transformed into
(sometimes called the “inverse Simpson concentration”). Second, the Shannon entropy
gets transformed into
Actually, MacArthur and Whittaker both argued long ago that when it comes to measuring diversity, the exponential of Shannon entropy is more natural than Shannon entropy itself.
Jost wasn’t the first person to argue for the use of effective numbers, but either he argued more convincingly or the time was ripe, because that view seems to have gone from controversial to commonplace extremely quickly. See for example the editorial introduction by Ellison to the special section on diversity in Vol. 91 of Ecology.
Jane wrote:
I haven’t thought much about complexity and don’t have well-developed opinions on how the word should be used. But I very much approve of being precise about the meanings of words such as “complexity” and “diversity”.
Part of the problem with “diversity” is that people have wanted to use it to mean too many things. Lou Jost has written vividly about this:
Eugene: that’s a good question. I’m not sure that the word “diversity” should reflect interactions between species. But the measures described above can be used in a sensible way to measure the amount of interaction between species.
(Actually, it’s an inverse relationship: the more interaction between species, the lower this diversity value is going to be, as we’ll see.)
Let’s take the simplest possible scenario: given a pair or species, either they’re deemed to interact, or they’re not. So we have an (unoriented) graph whose vertices are the species, and with an edge between species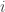 and
and 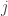 if and only if those two species interact.
if and only if those two species interact.
This is, equivalently, a symmetric square matrix of 0s and 1s. Call it . As usual, I’ll write
. As usual, I’ll write  for the proportions in which the species are present.
for the proportions in which the species are present.
Is the diversity in any way interesting?
in any way interesting?
Yes, as it turns out. Take your favourite value of , then choose the value of
, then choose the value of  that maximizes the diversity of order
that maximizes the diversity of order  . That maximum diversity value turns out to be equal to the largest number of non-interacting species. In other words, it’s the largest
. That maximum diversity value turns out to be equal to the largest number of non-interacting species. In other words, it’s the largest 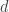 such that there exist
such that there exist 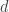 species none of which interact with any of the others. This is true independently of
species none of which interact with any of the others. This is true independently of  .
.
I don’t know of any connection with simplicial sets. Maybe they’d come in if you were thinking of multiple species interacting at once, rather than just interacting in pairs. So the graph I just described would turn into a simplicial complex. I have no idea what happens then.
Tom: First, an apology — I am a bit stymied by not knowing how to post links to this blog.
A few years ago I got intrigued by the so called “groupoid formalism” of Golubitsky, Stewart and their collaborators for the so called “coupled cell networks” ( a survey paper is here:
http://dx.doi.org/10.1090/S0273-0979-06-01108-6 ). I wanted to know what the groupoids in question were acting on.
This is how I got sucked into the networks business. For me to understand what Golubitsky et alii have done requires a doze of category theory. Now, there are two other efforts ( that I know of) to use category theory to understand networks — John’s project on this blog and David Spivak’s categorical informatics: http://math.mit.edu/~dspivak/informatics/
It feels like one should meaningfully model an ecosystem as a network. I am not sure if one should model such a network as a undirected graph, a directed graph, a Petri net or some sort of a higher dimensional model as in
Click to access HDMSN.pdf
(the latter paper is the reason why I mentioned simplicial sets).
But if one models an ecosystem as a directed graph,
one could try to understand and measure its complexity. The measures of diversity you nicely described seem to capture the diversity of nodes of such graphs. I am wondering if one could get at the diversity of edges or maybe higher dimensional versions of edges.
Admittedly this is a bit of a confused comment. But I have to run.
Hi, Eugene! I hope you had a nice jog.
Maybe you meant post links on this blog? For that, you can either just type in URLs as you did, or (for a superficial appearance of sophistication) use html commands, written in lower-case letters. For example, typing this:
• John Baez, <a href = “http://math.ucr.edu/home/baez/renyi.pdf”>Renyi entropy and free energy</a>.
will produce a link like this:
• John Baez, Renyi entropy and free energy.
But by the way: if you want to link to a specific comment on this blog, just click on the date of that entry, then look up at the little window on your browser that shows URLs, and you’ll see the URL of that comment, which you can use to link to it.
Maybe you meant a dose of category theory, but I kind of like this. A pride of lions, an exultation of larks, a doze of category theory.
“A doze of category theory” will live in cyberspaces forever.
I also don’t know how to put food on my family.
A speculative remark:
In theoretical physics, quantum field theory or statistical mechanics, a technique known as renormalization is often used. The renormalization determines how, for a given theory, evolve the parameters of this theory, from “large” scale to “small” scale. For instance, the scale may be the energy, and the parameters may be the charge or masses of particles.
Applied to biodiversity, “small” scale, could mean, for example, genus, and “large” scale could mean species. Of course, a hierarchy can be more elaborate than species / genus, but the idea of scale could be still valid.
Then we can study a problem of biodiversity at a specific scale. The result depends on the scale that is used. And maybe, we can relate the biodiversity at one scale to the biodiversity at another scale, and maybe find some kind of similarity with renormalization techniques.
I don’t know anything about renormalization in physics, but yes, one can think about varying the “scale” when assessing biodiversity.
Suppose we want to quantify the genetic diversity of a bird community. An initial, rough and ready survey records only the various genera and their abundances; similarities between genera are given by an average genetic distance. A second, more detailed survey records the relative abundances of, and genetic distances between, species. In principle, there might even be a third survey recording the genetic distances between individuals.
After each of the three surveys we use the abundance and distance data to calculate the diversity, giving values . Using the measures that Christina and I propose, the progression of diversity values reflects exactly the knowledge gained by the successively more refined surveys.
. Using the measures that Christina and I propose, the progression of diversity values reflects exactly the knowledge gained by the successively more refined surveys.
For instance, in the unlikely event that every species consisted of genetically identical individuals, we would have . (In our paper, we call this the “identical species” property.) A small amount of intraspecific variation would cause a small difference between
. (In our paper, we call this the “identical species” property.) A small amount of intraspecific variation would cause a small difference between  and
and  ; a moderate amount would cause a moderate difference; and so on.
; a moderate amount would cause a moderate difference; and so on.
Tom: do you or Cristina have any plans to go further with this? For example: any plans to dive into that ‘world of uncertainty’? I recently met someone named Jochen Jaeger, who came to Singapore for a workshop where they were developing an index of biodiversity for cities. These things can have real policy implications…
John, yes, Christina and I do have plans to go further with this. In particular, as you know, we’re planning in particular to go further with it at the conference and research programme in Barcelona next summer.
You asked about the “world of uncertainty” that opens up when you try to quantify the similarity between two species. (The diversity measures defined above require choosing a number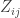 between 0 and 1 for each pair
between 0 and 1 for each pair 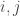 of species.) So let me say something about that.
of species.) So let me say something about that.
My overall view is that it’s up to the experts in particular fields should decide what measure of similarity is most meaningful.
For example, it’s easy to understand “genetic similarity” in a simplistic sense as “proportion of places where the DNA agrees”: e.g. the similarity between GTTAC and ACTGC would be 0.4, because they agree in 2 out of 5 places.
But after a moment’s thought you realize that it’s going to be way more complicated than that. DNA varies from individual to individual, so whose DNA do we use? And portions of the sequence of nucleotides (the letters G, T, A and C) are often repeated – e.g. ACTGC might mutate into ACTTGC – so you don’t really just want to put two DNA molecules next to each other and compare them position by position.
So there are subtleties here, and it might be that no way of measuring genetic similarity is clearly superior to all others. The same will probably be true for other, non-genetic, ways of measuring similarity between species. It’s definitely not for us to impose our opinion here.
The bottom line is that different notions of similarity between species give rise to different notions of diversity. That’s as it should be.
To take a rather artificial example, suppose you’re interested in the animals that live in a certain cave system. Perhaps you’re an expert on animal locomotion, and you’re studying how the animals get around these caves. Some crawl, some swim and some fly. From that point of view, bats and birds would be deemed to be quite similar. But phylogenetically they’re very different: they diverged a long time ago in their evolution.
So if the caves largely consisted of flying creatures, an animal locomotion expert might want to view the community as not very diverse, but a phylogeneticist might “disagree”. I put it in quotation marks because these two people don’t really disagree — they’re just measuring different things.
All that should be kind of obvious. But it seems there’s a necessity to be clear about it. There’s a lot of discussion at the moment about how the word “diversity” should be used.
When machine learning folk look for a similarity measure on strings, they often choose a string kernel, which allows for non-contiguous sequence matching.
Thanks, David. I hope you didn’t take me to be suggesting that this problem of comparing nucleotide sequences was unsolved. I know for sure that the people who do these things have their ways. I only wanted to point out that even once you’ve made a broad choice such as “I want my measure of similarity to be genetic”, there are still a bunch of finer choices to be made.
Different choices embody different views on what it means for two species to be similar, and therefore lead to different measures of a community’s diversity.
This is just a quick thought as I walk to the train…
It almost seems like diversity is not zero dimensional, but should take into consideration the full curve.
Sorry. I didn’t say that quite right. What I meant is maybe diversity should be thought of in terms of the whole curve and not just a single real number.
No I didn’t think you were suggesting that. I was recalling the fun we once had with kernels. There’s been lots of work done on phylogenetic tree reconstruction using kernel methods.
Eric wrote:
Yes, absolutely. That’s one of the points we’re keen to make.
In fact, it can be shown that in a certain precise sense, the relative abundances can be reconstructed from the curve. So the process of passing from the relative abundance vector to the curve doesn’t lose any information: it’s just repackaging it in a useful way.
can be reconstructed from the curve. So the process of passing from the relative abundance vector to the curve doesn’t lose any information: it’s just repackaging it in a useful way.
(I feel like there should be some integral transform in the background here.)
David: OK, good. Thanks. My colleague Chris Quince in the engineering department works on things like the human microbiome — the enormous community of microorganisms living in and on your body (and apparently outnumbering your actually body cells by 10:1). He’s explained to me that to calculate the diversity of microbial samples, they extract some DNA and use some rather sophisticated algorithms to reconstruct something like a phylogenetic tree. It seems to be quite complex stuff, not necessarily conceptually right, but somehow accepted by that community.
Oops: when I wrote that the algorithm was “accepted by that community”, I didn’t mean the community of microbes living in and on your body. I meant that community of researchers.
In further response to John’s comment, here’s a question I’ve been thinking about on and off: how should we understand non-symmetric similarity matrices?
For a similarity matrix to be symmetric means that the similarity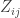 between the
between the 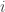 th and
th and 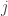 th species is equal to the similarity
th species is equal to the similarity  between the
between the 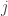 th and
th and 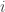 th species. This sounds like an obvious assumption. However, none of the properties of diversity that we prove in our paper turn out to require it.
th species. This sounds like an obvious assumption. However, none of the properties of diversity that we prove in our paper turn out to require it.
Moreover, there are interesting and useful non-symmetric similarity matrices. For example, in the appendix to our paper, we show that a certain phylogenetic diversity measure (proposed by Chao, Chiu and Jost) can be obtained from ours by sticking in a certain similarity matrix . And this matrix
. And this matrix  is non-symmetric.
is non-symmetric.
In that situation, the lack of symmetry comes from the fact that time is not symmetric. Saying that species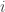 is descended from species
is descended from species 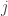 , is not the same as saying that species
, is not the same as saying that species 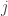 is descended from species
is descended from species 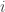 . Phylogenetic trees have a definite orientation.
. Phylogenetic trees have a definite orientation.
There’s at least one other way in which non-symmetric similarity matrices arise naturally: disease transmission networks.
(It’s not clear that it’s appropriate to use the word “diversity” here, but our measures do seem to measure something useful.)
do seem to measure something useful.)
Suppose we have a group of individuals, each of whom may or may not be infected with a certain disease. For each pair
individuals, each of whom may or may not be infected with a certain disease. For each pair  of individuals, we can guess the probability
of individuals, we can guess the probability 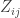 of the
of the 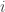 th individual passing the disease to the
th individual passing the disease to the 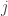 th individual within a one-year period. Since our “units” are not species but individuals, we should take our
th individual within a one-year period. Since our “units” are not species but individuals, we should take our 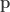 (which would ordinarily specify the proportions in which the species are present) to be
(which would ordinarily specify the proportions in which the species are present) to be 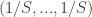 .
.
The “diversity” measures something like the robustness of the group. A high value means that it takes a long time for a disease to spread. There is the idea that diverse populations are less vulnerable to disease, but I’m not actually saying anything about that: I’m just saying what the quantity that we called “diversity” measures.
measures something like the robustness of the group. A high value means that it takes a long time for a disease to spread. There is the idea that diverse populations are less vulnerable to disease, but I’m not actually saying anything about that: I’m just saying what the quantity that we called “diversity” measures.
However, it’s entirely possible that the similarity matrix is non-symmetric. For example, some sexually transmitted infections are more easily passed from men to women. Or the individuals might be of different species, etc. So what we have here is a useful but non-symmetric similarity matrix.
is non-symmetric. For example, some sexually transmitted infections are more easily passed from men to women. Or the individuals might be of different species, etc. So what we have here is a useful but non-symmetric similarity matrix.
Very nice article, also the series at the n-café (which is probably the only one I understand there). Understanding diversity is hugely important giving the rate at which we are destroying it (at least, for many of it’s possible meanings).
The following is primarily a problem for ecologists rather than mathematicians, but which connection are there between ecosystem functioning (and the services it may provide) and the diversity it contains?
I’m perfectly aware that if I want a documented answer (or some collection of them) I should read into the extensive research over this particular question. (And indeed tomorrow I’ll borrow the book “Biodiversity and ecosystem functioning” from my faculty’s library). But maybe some of the writers or readers of this blog have a different background and so novel perspectives particularly to the way biodiversity is proposed to be measured here (for example, with diversity profiles).
I have my own intuitions on the subject but wouldn’t like yet to influence others.
Baez, when you said:
Well, looking for those appropriate constructs are my favourite mathematical activity. One I’ve learnt from your posts is the n-cube graph as, for example, a model of the configuration space for an ecological succession.
Thanks very much, Matías.
I know that the question you ask — the connection between diversity and ecosystem functioning — has been extensively considered, but I know very little about it.
Just about the only thing I think I know is that it’s the high values of that are generally considered to be most important for the functioning of whole ecosystems. The general thought, crudely put, is that common species affect the functioning of an ecosystem more than rare ones. And high values of
that are generally considered to be most important for the functioning of whole ecosystems. The general thought, crudely put, is that common species affect the functioning of an ecosystem more than rare ones. And high values of  pay more attention to common species than low values do.
pay more attention to common species than low values do.
Take, for example, the traditional or “naive” model in which the similarities between species are ignored. Then the diversity of order is the so-called Berger–Parker index. This is defined as
is the so-called Berger–Parker index. This is defined as
In other words, you find the most common species in the community or ecosystem, then calculate the reciprocal of its relative abundance.
As Berger and Parker pointed out (in a study of plankton sediment on the sea floor), it’s a measure of dominance. A high Berger–Parker index means that the most common species isn’t very common: no single species is too dominant. A low value means that some particular species makes up a large proportion of the population. For example, a Berger–Parker index of 1.25 = 5/4 means that 4/5 of your community comes from one particular species.
The Berger–Parker index is a special case of our measures: it’s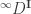 , where
, where  is the identity matrix. (The similarity matrix is
is the identity matrix. (The similarity matrix is  because we’re using this naive model.) So it’s an effective number. This means that it’s measured in units of species: a Berger–Parker index of 1.25 means that your community “effectively contains 1.25 species”.
because we’re using this naive model.) So it’s an effective number. This means that it’s measured in units of species: a Berger–Parker index of 1.25 means that your community “effectively contains 1.25 species”.
That’s an important point. Reducing a distribution to a single number can be useful, but it’s always risky.
My guess is that you meant to attach this comment as a reply to this one, where Eric was suggesting that we use the diversities for all instead of just one. This is mathematically very similar to using the whole partition function to characterize a physical system, instead of its value at a single temperature.
instead of just one. This is mathematically very similar to using the whole partition function to characterize a physical system, instead of its value at a single temperature.
I’m sure most people here know what I’ll say in the next paragraph, but I can’t resist saying it anyway:
The main advantage of using just one real number to measure how good things are, instead of something more complicated, is that real numbers are linearly ordered: either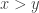 or
or 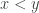 or
or 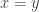 . So, if we’re trying to make decisions about what’s ‘best’, measuring ‘goodness’ by a single real number lets us decide this unambiguously. However, there’s almost always a huge price: if we don’t make the best decision about how to measure goodness by single real number, what’s ‘best’ won’t really be best!
. So, if we’re trying to make decisions about what’s ‘best’, measuring ‘goodness’ by a single real number lets us decide this unambiguously. However, there’s almost always a huge price: if we don’t make the best decision about how to measure goodness by single real number, what’s ‘best’ won’t really be best!
In particular—and this may be more controversial—I think that the idea in economics of assigning a numerical value to happiness (or more precisely ‘utility’) is misguided, because in real life the conditions of the von Neumann–Morgenstern utility theorem rarely hold. People don’t actually make decisions by comparing all possible options and ranking them! Often the choice I would have made, if I’d thought of it, is one I never considered. But the idea of utility offers a nice justification for the existence of money, and money is convenient, so the idea of utility will be with us for a long time.
Yep. :-) The system wouldn’t let me reply directly to Eric’s post.
It may be useful to note that biologists will often reduce their data to a single number (an index, a mean, whatever) without any thought about what is being lost. If the folks commenting on this blog need to be reminded of the usefulness of single numbers, it’s the opposite for many biologists.
Jane wrote:
Right. The column of text gets a bit more skinny with each successive reply to a reply to a reply… and after 3 levels of reply, it becomes absurdly skinny, so the system doesn’t let you reply directly. In that situation you should go back up to the first reply you can comment on.
In fact this what Eric had to do to write the comment you were trying to reply to! You can see this by looking at the comment he was replying to: it has no blue Reply thing under it, unlike the comment you just wrote, or the one here.
Indeed. People who don’t like math much tend to use numbers where more sophisticated constructs might be better. These are often the people who think math is about numbers.
Digressing a bit… I read that the dean of admissions at MIT recently said:
And how do we improve the gender balance in math? Get the story out that it’s not about sitting a desk doing science and engineering all day?
Yes, it annoys me when my algebra students, say, make an error in elementary arithmetic and say “My only problem was the math.”. What’s ironic is that they usually don’t realise how true that is!
Jane, yes, I agree. In her 1975 book Ecological Diversity, Evelyn Pielou warned that
All locallly adapted diversities follow this PDF for relative abundance diversity (RAD):
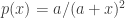
If it is a meta-community then the RAD is spread out by the log of the two ratios.
[…] in a historically accurate way, Computational Complexity gave a short introduction to bitcoins, Azimuth had a guest post on measuring bio diversity, and Casting Out Nines explains that math is not too […]
[…] In biology, entropy is one of many ways people measure biodiversity. For a quick intro to some of the issues involved, try: Tom Leinster, Measuring biodiversity, Azimuth, 7 November 2011 […]